 
80S ribosomes and 40S subunits are depicted as silhouettes of the crystal structures of bacterial 70S and 30S ribosomal species, with approximate locations of the aminoacyl-tRNA (A), peptidyl-tRNA (P), and exit (E) sites labeled in the 40S subunit. The process of initiation is depicted as a pathway of reactions (a subset described in blue type), beginning with the dissociation of 80S ribosomes into free 40S and 60S subunits and the assembly of the 43S PIC on the small ribosomal subunit. Pathway of eukaryotic translation initiation via ribosomal scanning. It has also emerged that multiple DEAD-box RNA helicases participate in producing a single-stranded "landing pad" for the 40S subunit and in removing the secondary structure to enable the mRNA to traverse the 40S mRNA-binding channel in the single-stranded form for base-by-base inspection in the P site. A robust molecular model now exists, describing the roles of initiation factors, notably eukaryotic initiation factor 1 (eIF1) and eIF1A, in stabilizing an "open" conformation of the 40S subunit with Met-tRNAiMet bound in a low-affinity state conducive to scanning and in triggering rearrangement into a "closed" conformation incompatible with scanning, which features Met-tRNAiMet more tightly bound to the "P" site and base paired with AUG. Over the past decade, a combination of yeast genetics, biochemical analysis in reconstituted systems, and structural biology has enabled great progress in deciphering the mechanism of ribosomal scanning. 
For most mRNAs in eukaryotic cells, this is accomplished by the scanning mechanism, wherein the small (40S) ribosomal subunit attaches to the 5' end of the mRNA and then inspects the leader base by base for an AUG in a suitable context, using complementarity with the anticodon of methionyl initiator tRNA (Met-tRNAiMet) as the key means of identifying AUG. Testing the fluorescence of E.The correct translation of mRNA depends critically on the ability to initiate at the right AUG codon. 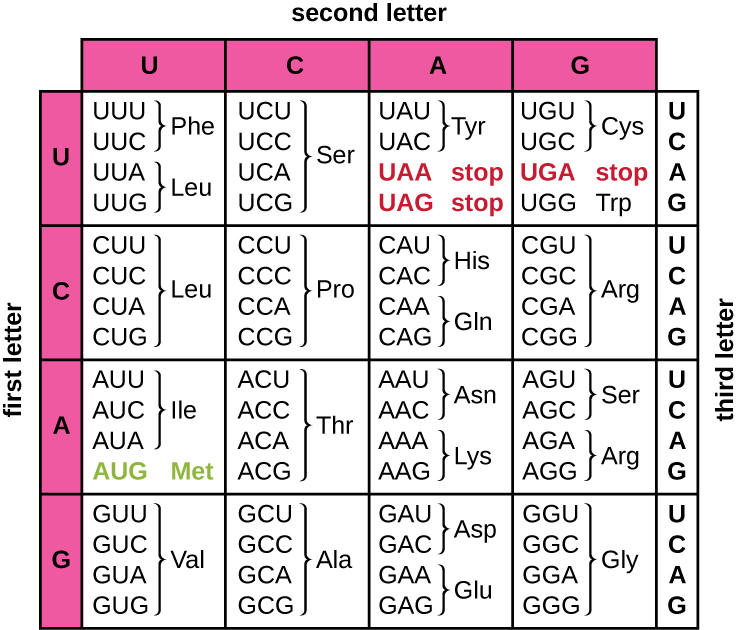
A plasmid has been engineered with two start codons slightly out of frame from each other, but close enough to test start codon slippage. This construct aims to test the possibility of multiple start codons on a single mRNA strand. Which frame will be preferred? The Objective We are looking into what happens when you have multiple start codons close together, but in different reading frames. If you advance it again you are now back in your first reading frame, but you have skipped the first codon. If you advance you starting point one more base, that is the third reading frame. If you then move your start point ahead one base, then you are reading in a new reading frame. If you start at one base and read the bases in groups of three from that point on, that is one frame of reference. Remember those codons and how they are groups of three bases on the mRNA? The reading frame refers to which group of three. Out of frame refers to how the ribosome reads the mRNA. We are trying to understand what happens when there are two start codons very close together, but out of frame. But what happens if you have two AUG codons close together? That is the question we are attempting to answer. Once this start codon in read the ribosome will continue reading and building the polypeptide (protein) until a stop codon is reached. 
This start codon is generally AUG, or Methionine. Instead, once the mRNA is bound by a ribosome, a start codon must first be read before the protein can be translated. However, much like the lines on a sheet of paper, the protein coding sequence of an mRNA molecule does not start right at the beginning, or top of the page. MRNA is the molecule that carries information about the sequence of amino acids in a protein. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
